What is Point Mutation?
A point mutation is a change in one base pair in the nucleotide sequence of the DNA strand.
It can be due to an error during DNA replication but can occur as a result of exposure to radiation, such as X-rays.
There are two main types of point mutations:
- Transition mutations
- Transversion mutations
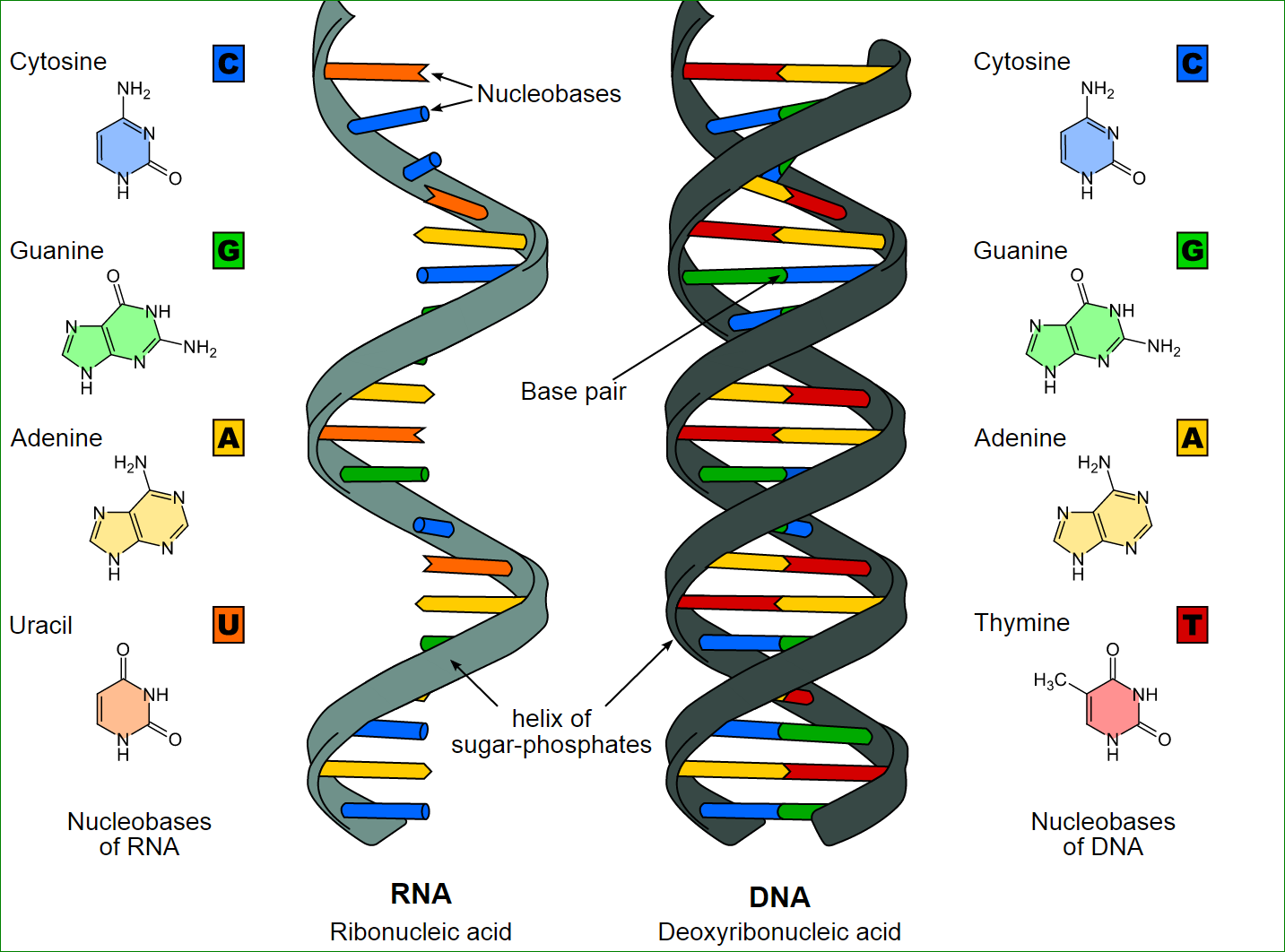
On the double-stranded DNA, pyrimidine bases bond with purine bases.Thymine and cytosine are pyrimidine bases, while adenine and guanine are purine bases; and thymine bonds with adenine while cytosine bonds with guanine.
In a transition mutation, there is a substitution of one pyrimidine base for another, for example cytosine could be substituted for thymine, or vice versa.
Similarly in a transition mutation, a purine base can substitute for another purine base, so guanine could substitute for cytosine or vice versa.
Here is an example:
If we have a sequence of DNA: TAC GAA TCA GCT
Transcribed mRNA: AUG CUU AGU CGA
Translated into amino acid: Methionine Leucine Serine Arginine
With a substitution of a purine base, so say base C is replaced by G so GCT in the DNA becomes GGT.
Our new sequence of DNA: TAC GAA TCA GGT
New transcribed mRNA: AUG CUU AGU CCA
Translated into amino acid: Methionine Leucine Serine Proline
Notice, that only one amino acid has changed here, so instead of Arginine we form the amino acid Proline. However, the other amino acids remain the same. The result could be a silent mutation where even though there is a change in the amino acid, the protein is unaffected. Of course it could result in a missense mutation where the protein is adversely affected. This could result in disease.
In a transversion mutation a pyrimidine base substitutes for a purine base, or a purine base substitutes for a pyrimidine base.
An example of a disease caused by a point mutation is sickle-cell anemia.
What happens in sickle-cell is that the mutation results in valine being formed instead of glutamic acid causing a change in the structure of the hemoglobin.
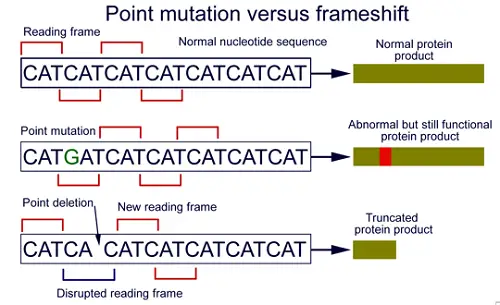
What is Frameshift Mutation?
A frameshift mutation is a change in the genetic code in which a nucleotide base is either:
- added to, inserted into the sequence of nucleotides, or
- removed, deleted from the sequence of nucleotides.
Since translation and subsequent protein synthesis relies on reading codons in triplets the result is an incorrect message being translated.
The reason this mutation is called a frameshift mutation is that it literally results in a shift of the reading frame. This means that if the frames are wrong, then each read will be wrong since there will be either one extra base or one less base in the sequence.
More than one insertion or deletion can occur which can still produce a viable protein. This may then result in a different variant which may or may not be advantageous to the individual.
The reading frame in genetics refers to the codons which are arranged in non-overlapping threes, triplets. Each triplet codes for a specific amino acid.
The codons are the bases on the DNA or mRNA molecule.
Here is an example:
If we have a sequence of DNA: TAC GAA TCA GCT
Transcribed mRNA: AUG CUU AGU CGA
Translated into amino acid: Methionine Leucine Serine Arginine
With an insertion of a base before the T in TAC:
The sequence of DNA: ATA CGA ATC AGC T
NOW the transcribed mRNA becomes: UAU GCU UAG UCG A
Amino acid that would result: Tyrosine Alanine STOP Serine
Notice the frame has shifted down by one base. This changes the entire amino acid sequence from that point onwards. What is even worse is that in this hypothetical example, we now have a STOP codon instead of Serine, so translation halts. Thus protein synthesis stops and the Serine is not even produced.
The frameshift mutations can produce a strand that is either too short because of deletion of a base, or too long due to insertion of a nucleotide base.
Frameshift mutations are responsible for some very severe diseases, for instance, Tay-Sachs disease.
In Tay Sachs disease, a mutation occurs causing a change in an enzyme.
What is the difference between Point and Frameshift Mutation
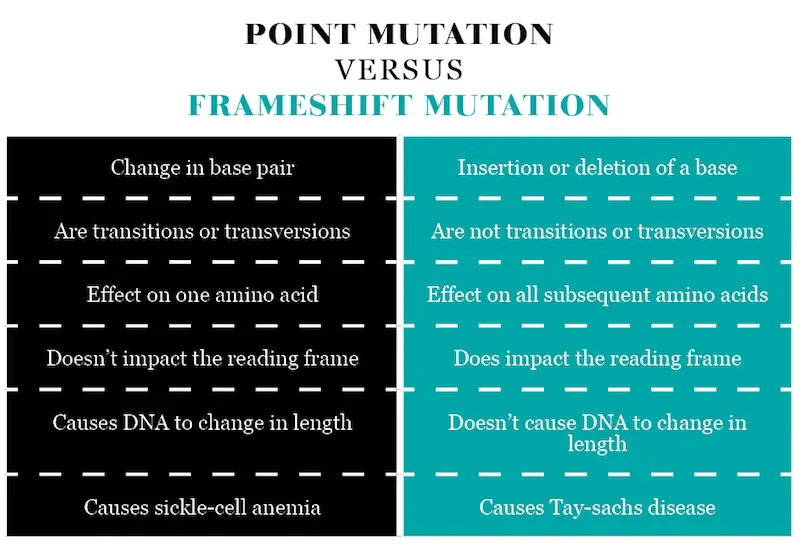
- A point mutation is a change in a base pair, while a frameshift mutation is a base insertion or base deletion.
- Point mutations are transitions or transversions, frameshift mutations are not.
- A single point mutation only affects one amino acid, while a single frameshift mutation affects the entire subsequent amino acids.
- Point mutations don’t impact the reading frame while frameshift mutations do.
- Point mutations cause the DNA strand to change in length while frameshift mutations don’t cause this.
- A disease caused by a point mutation is sickle-cell anemia, while a disease caused by a frameshift mutation is Tay-Sachs disease.
Table comparing Point and Frameshift Mutation

Summary:
- Point mutations cause a change in nucleotide base pairs. These can be transitions or transversions.
- Frameshift mutations cause a change by either an insertion of a base or a deletion of a base in DNA or mRNA.
- Point mutations do not impact the reading frame of the strand since they only impact the translation of one amino acid where they occur.
- Frameshift mutations impact all subsequent amino acids because they cause a shift in the frame, because all the triplet codons change subsequent to the location of the mutation.
- These mutations can be caused by errors during DNA replication or due to exposure to radiation.
- Both these mutations can cause severe diseases such as Tay-sachs and sickle-cell disease.
Author: Dr. Rae Osborn
Dr. Rae Osborn holds Honours Bachelor of Science degrees in Zoology and Entomology, and Masters of Science in Entomology from the University of Natal in South Africa. She has received a PhD in Quantitative Biology from the University of Texas at Arlington. She was a tenured Associate Professor of Biology at Northwestern State University in Louisiana for 10 years. She also completed an AAS Degree in Information Network Specialist and an AAS in Computer Information Systems, at Bossier Parish Community College in Louisiana.









Leave a Reply